 Although single cell RNA sequencing studies have begun
providing compendia of cell expression profiles, it has
proven more difficult to systematically identify and
localize all molecular cell types in individual organs to
create a full molecular cell atlas. From droplet- and
plate-based single cell RNA sequencing applied to ~75,000
human lung and blood cells, combined with a multi-pronged
cell annotation approach that includes extensive tissue
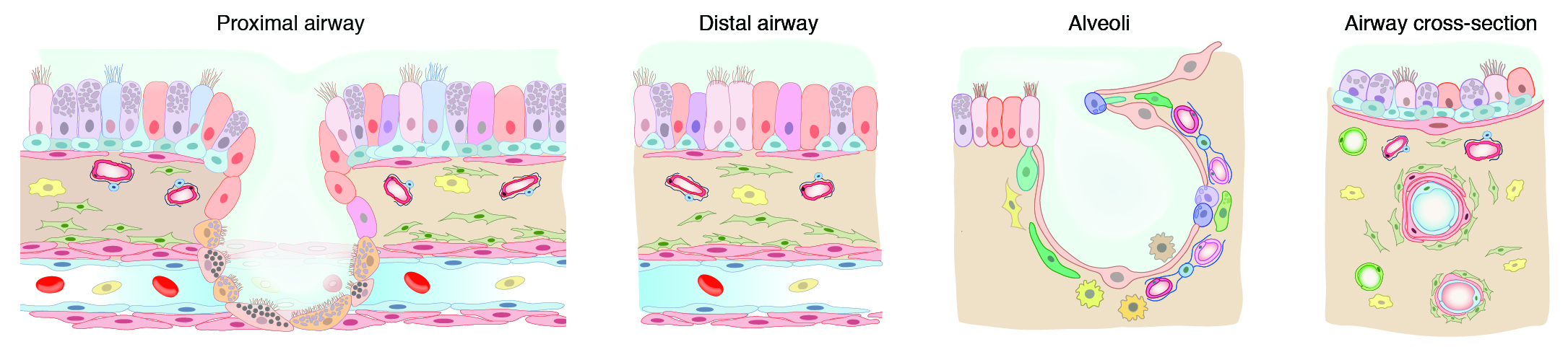
localization, we have defined the gene expression profiles
and anatomical locations of 58 cell populations in the human
lung, including 41 of 45 previously known cell types or
subtypes and 14 new ones. Learn more in our
manuscript
and our GitHub
repository.
Although single cell RNA sequencing studies have begun
providing compendia of cell expression profiles, it has
proven more difficult to systematically identify and
localize all molecular cell types in individual organs to
create a full molecular cell atlas. From droplet- and
plate-based single cell RNA sequencing applied to ~75,000
human lung and blood cells, combined with a multi-pronged
cell annotation approach that includes extensive tissue
localization, we have defined the gene expression profiles
and anatomical locations of 58 cell populations in the human
lung, including 41 of 45 previously known cell types or
subtypes and 14 new ones. Learn more in our
manuscript
and our GitHub
repository.
cellxgene
Explore the data in your browser on cellxgene.cziscience.com hosted by the Chan Zuckerberg Initiative:
Data availability
Thank you for your interest in obtaining access to Dr. Krasnow’s and Dr. Quake's lung single-cell RNA sequencing datasets. De-identified, non-raw data (count/UMI tables, metadata, and Seurat and scanpy objects for your own bioinformatic piplines) can be downloaded directly from Synapse. Because of human patient privacy considerations, in order to access raw sequencing reads, you must first establish an account with the European Genome-phenome Archive (EGA), submit a request to access EGAC00001001536, and sign a Data Access Agreement (DAA). You will receive a separate email from the Data Access Committee (DAC) containing additional instructions once your EGA request has been submitted.

